In silico Antibody-Peptide Epitope prediction for Personalized cancer therapy
The human leukocyte antigen (HLA) system is a complex of genes on chromosome 6 in humans that encodes cell-surface proteins responsible for regulating the immune system. Viral peptides presented to cancer cell surfaces by the HLA trigger the immune system to kill the cells, creating Antibody-peptide epitopes (APE). This study proposes an in-silico approach to identify patient-specific APEs by applying complex networks diagnostics on a novel multiplex data structure as input for a deep learning model. The proposed analytical model identifies patient and tumor-specific APEs with as few as 20 labeled data points.
Additionally, the proposed data structure employs complex network theory and other statistical approaches that can better explain and reduce the black box effect of deep learning. The proposed approach achieves an F1-score of 80\% and 93\% on patients one and two respectively and above 90\% on tumor-specific tasks. Additionally, it minimizes the required training time and the number of parameters.
The human leukocyte antigen (HLA) system or complex is a complex of genes on chromosome 6 in humans that encode cell-surface proteins responsible for regulating the immune system. The HLA system also known as the human version of the major histocompatibility complex (MHC) is found in many animals.
HLA genes are highly polymorphic, which means that there are thousands of different forms of these genes called alleles, allowing them to fine-tune the adaptive immune system. The proteins encoded by certain genes are also known as antigens, because of their historic discovery as factors in organ transplants.
As shown in Figure \ref{fig:1} HLA’s proteins present viral peptides from inside the cell to the surface of the cell. For example, if the cell is infected by a virus or is cancerous, the HLA system brings abnormal fragments, called peptides, to the surface of the cell so that the cell can be destroyed by the immune system.
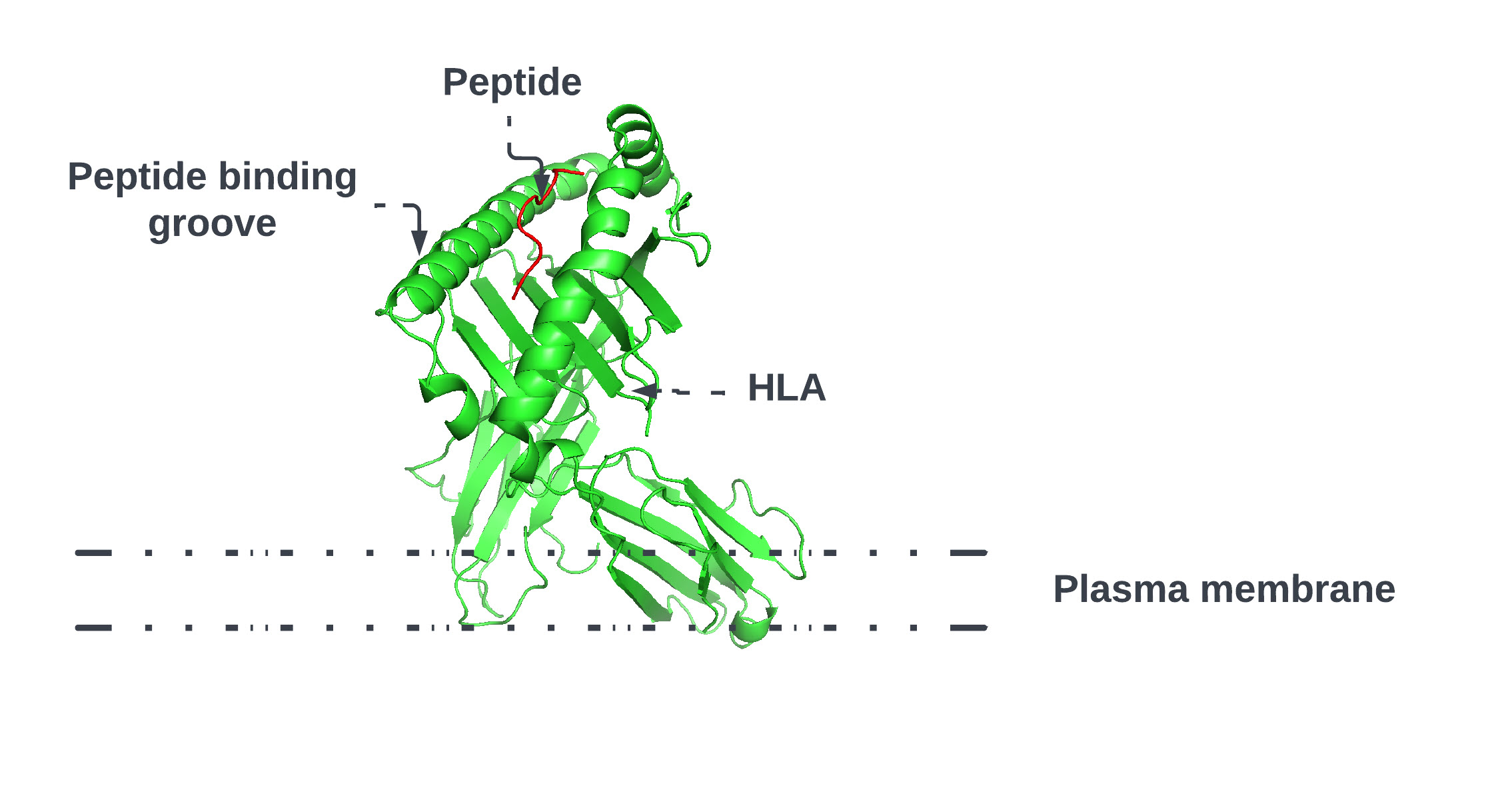
Predicting the specific HLA peptide combination that will present the peptide to the cell’s surface permits the creation of a treatment that will trigger the human immune system to destroy the cell.
Specifically, in cancer, this ability is essential, given that cancer is highly mutagenic with tumor and patient-specific mutations. This means that patients with the same tumor type will have different mutations that result in different reactions to the same treatment.
Advances in Deoxyribonucleic acid (DNA) sequencing, Messenger Ribonucleic acid (mRNA) vaccines, and high computational power allow us to work toward patient-specific therapy. This approach, called personalized mRNA-based antitumor vaccine, visualized in Figure \ref{fig:2}, is bound to play a major role in the future.
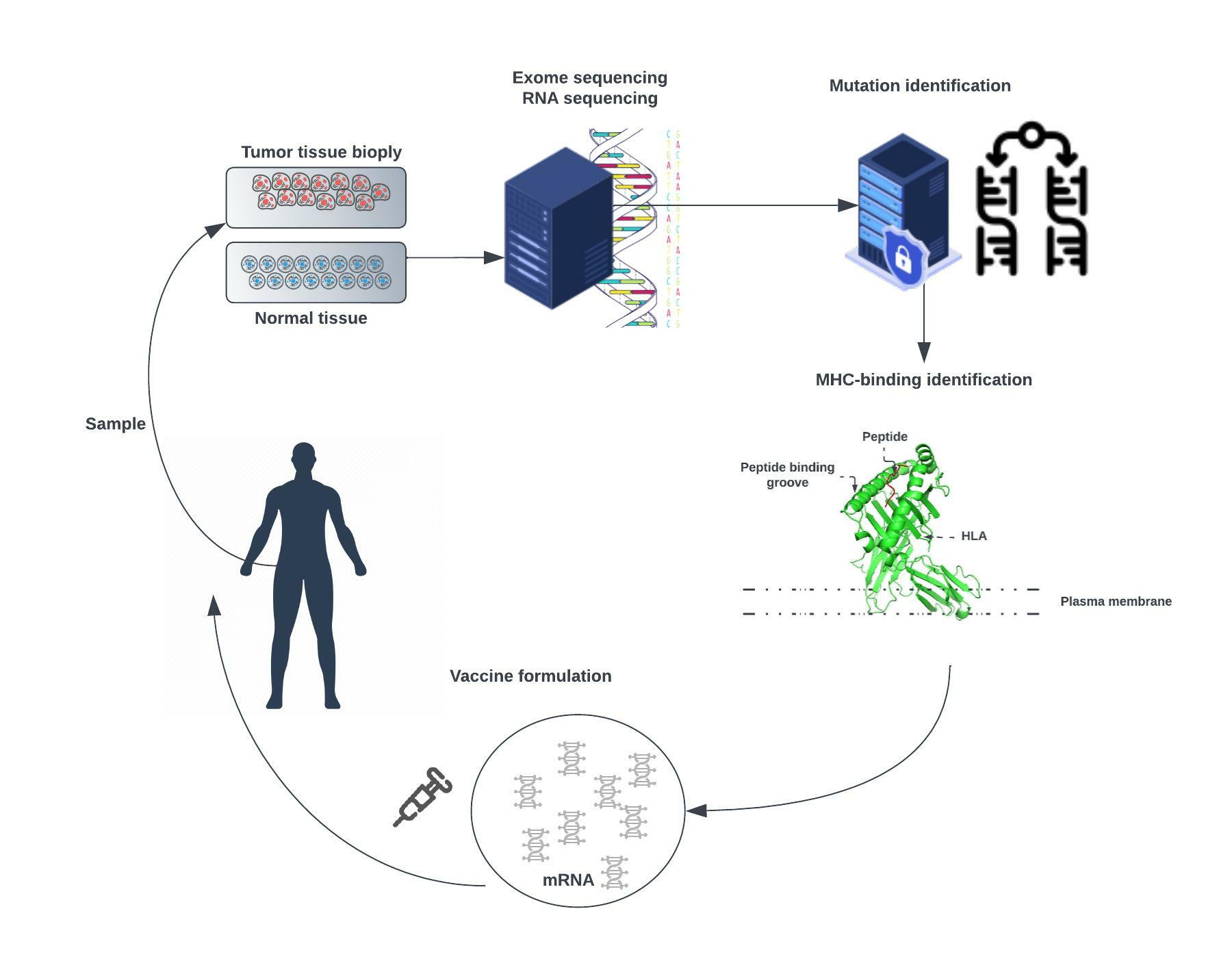
The approach is meant to trigger an antitumor immune response in patients by challenging them with mRNAs encoding tumor-specific antigens \citep{pastor_rna_2018}. These mRNAs can be directly injected as naked RNA or loaded into patient-derived dendritic cells.
In this work, we propose to extend the approach with additional laboratory and analytical optimization steps.
Concretely DNA sequenced from the patient is used to select candidate peptides that will result from gene expression. As the space of possible combinations is huge a subset of potential peptides is synthesized and their reaction to the patient’s specific HLA alleles is tested by applying an enzyme-linked immunospot (ELISpot) assay.
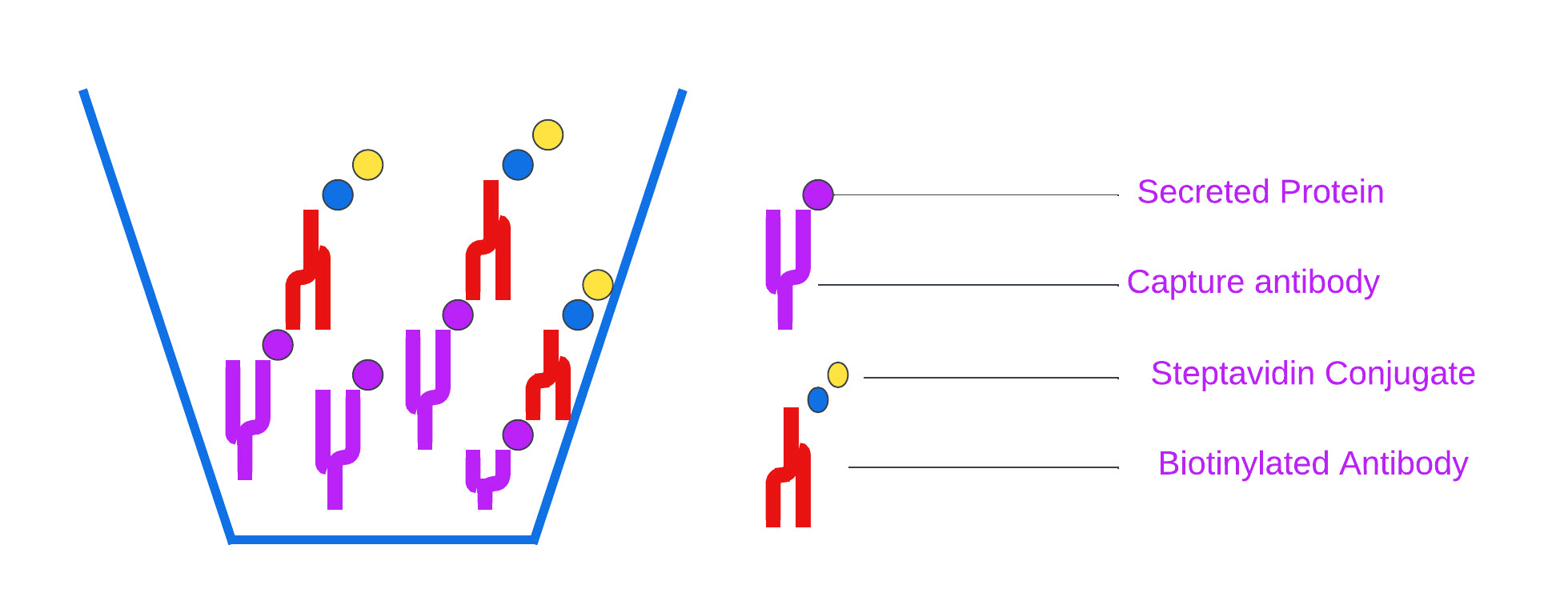
An enzyme-linked immunospot (ELISpot) assay \citep{engvall_enzyme-linked_1971}, shown in Figure \ref{fig:3}, is a highly versatile and sensitive technique that is used for qualitative and quantitative measurement of the cytokine-secreting cells at the single-cell level. \citep{paulie_chapter_2006}